·Experimental Research·
Network pharmacology analysis of ginkgo leaf extract against diabetic retinopathy
Bai Wen, Jiang Qin
Eye Hospital, Nanjing Medical University, Nanjing 210029, China
Corresponding author: Jiang Qin, Email: jqin710@vip.sina.com
[Abstract] [View PDF in English] [View PDF in Chinese] [Read Full Text]
Objective To identify the key targets of ginkgo leaf extract against diabetic retinopathy (DR), and to identify the underlying pharmacological mechanism using network pharmacology.
Methods Potential targets of active components in ginkgo leaves were searched from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform and SwissTargetPrediction database. Therapeutic targets highly related to DR were retrieved from the GeneCards and DisGENET databases. Their intersection targets were subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses to construct the active component-target network. Based on the degree of network nodes, key active components and targets were used for molecular docking verification.
Results A total of 27 active components and 34 potential therapeutic targets of ginkgo leaf extract in the treatment of DR were screened. GO enrichment analysis indicated that these therapeutic targets were mainly enriched in the processes of inflammation, oxidative stress, angiogenesis, and hypoxic damage. KEGG pathway enrichment analysis revealed that they were mainly enriched in the advanced glycation end product-receptors for advanced glycation end products, mitogen active protein kinase, phosphatidylinositol-3-kinase-protein kinase B, hypoxia inducible factor-1, tumor necrosis factor (TNF), and interleukin (IL)-17 signaling pathways. According to degree values calculated from the active component-target network, the top four key active components with higher degrees were quercetin, kaempferol, luteolin, and isorhamnetin, and the top eight key targets were serine/threonine protein kinase 1, vascular endothelial growth factor A, IL-6, TNF, nitric oxide synthase 3, peroxisome proliferator activated receptor-γ, IL-10, and matrix metalloproteinase-9. The binding activities between key active components and targets were good.
Conclusions A variety of active compounds contained in ginkgo leaf extract targeted therapeutic targets related to DR, and regulated multiple signal pathways, thereby exerting a therapeutic effect on DR.
[Key words] Network pharmacology; Molecular docking simulation; Ginkgo leaves; Diabetic retinopathy
Fund program: National Natural Science Foundation of China (82070983)
DOI: 10.3760/cma.j.cn115989-20210105-00010
Diabetic retinopathy (DR) is a microvascular complication of diabetes. It can cause blindness in people over 50 years of age 1. Simply controlling blood glucose is not an effective way to delay the occurrence and development of DR. At present, therapeutic methods, mostly used in the treatment of advanced DR, such as laser photocoagulation, intravitreal injection of vascular endothelial growth factor (VEGF) drugs, and vitrectomy, have been proven to effectively inhibit the progression of DR, but have also been shown to damage some tissues 2-3. Due to natural and safe characteristics, traditional Chinese herbal medicines used in DR prevention and control have attracted more attention 4. Ginkgo leaves have been used clinically for thousands of years; Ginkgo biloba preparations have been used in more than 130 countries. Because of its extensive use, well-known experts from Integrated Traditional Chinese and Western Medicine, and the Chinese Medical Doctor Association, have prepared the Chinese Expert Consensus on Clinical Application of Oral Ginkgo Biloba Preparations as a guideline for the clinical use of ginkgo leaves. The Ginkgo biloba preparation is recommended for patients with early DR 5. According to numerous basic and clinical studies, ginkgo leaf extracts can safely and effectively minimize retinal microvascular and nerve damage, and it can be used as an alternative therapy for the prevention and treatment of DR 6-9. Ginkgo leaves contain many pharmacological components, such as flavonoids (quercetin, kaempferol, luteolin, and isorhamnetin, etc.) and terpenoids (ginkgolide, etc.) 10-11. These components may play a positive role in improving blood microcirculation, regulating lipid metabolism, preventing blood clot formation, resisting oxidative stress, and reducing inflammation 12-14. However, the specific mechanism of its treatment of DR has not been fully clarified. Therefore, it is necessary to further identify the active components in ginkgo leaves to identify the pathways and mechanisms by which these components successfully treat DR. In addition, the network pharmacology method, used in the field of ophthalmology, can be used to study the underlying mechanism of drug action in vivo, and reveal the regulatory principles of small molecule drugs 15-17. This study therefore aimed to identify the underlying mechanism of Ginkgo leaf extract in the treatment of DR by using a network pharmacological approach, to provide new ideas for the treatment of DR.
1 Materials and Methods
1.1 Screening of active components and targets of ginkgo leaf extracts from databases
The active components of ginkgo leaves with the threshold of oral bioavailability ≥ 30% and drug-likeness ≥ 0.18 were searched using the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (https://old.tcmsp-e.com/tcmsp.php). The TCMSP and SwissTargetPrediction database (http://www.swisstargetprediction.ch/) were also searched for known targets and predicted targets of active components in ginkgo leaves, by setting the threshold probability > 0.3.
1.2 Screening DR-related targets from databases, and identification of potential therapeutic targets
The targets related to the treatment of DR were identified using the GeneCards (https://www.genecards.org/) and DisGENET (https://www.disgenet.org/home/) databases by retrieving the term: diabetic retinopathy. Targets highly related to DR were screened with relevance score > 5 and gene-disease association score > 0.1 as thresholds. By intersecting these two targets, potential therapeutic targets against DR were identified.
1.3 Construction of the ginkgo lead extract active components-target network, and analysis of key nodes
The target protein was imported into the STRING (https://string-db.org/) database after the target name was normalized in the Uniprot (https://www.uniprot.org/) database. The interactions of target proteins with high confidence > 0.7 were imported into Cytoscape 3.8.2 software to construct an active components-potential therapeutic target network of ginkgo leaf extracts. The key active components and key targets with high importance according to degree value parameters were screened using the NetworkAnalyzer plug-in in Cytoscape 3.8.2 software. The degree value is the number of interactions between nodes and other nodes, which represents the importance of nodes.
1.4 Functional enrichment analysis and pathway analysis for biological functions and the signaling pathways of targets
The potential therapeutic targets of ginkgo leaf extract against DR were subjected to functional enrichment analysis, pathway analysis, and visualization using R software (R-3.6.3; The R Foundation for Statistical Computing, Vienna, Austria). In this process, the org.hs.eg., the db package was used to convert the above target names into entrez identifications, and the clusterProfiler package was used to perform Gene Ontology (GO) for identifying biological functions (with P-value < 0.05 and Q-value < 0.05 as thresholds), as well as the Kyoto Encyclopedia of Genes and Genomes (KEGG) signal pathway enrichment analysis to identify potential therapeutic targets. Finally, the enrichplot package was used to visualize the top 10 GO entries and the top 20 signal pathways in three modules of biological process, cell composition, and molecular function, where they were sorted by the count value of the number of enriched genes.
1.5 Molecular docking verification of the binding effect of key active components and key targets of ginkgo leaf extracts
Mol2 files for key active components were downloaded from the Pubchem (https://pubchem.ncbi.nlm.nih.gov/) database, which were then converted into three-dimensional (3D) structure files in PDF using Chem3D software. For key target proteins, the 3D structure files in PDP format were directly downloaded from the Protein Database (http://www.rcsb.org/pdb). Autodock 4.2 software was then used to set Gridbox parameter ranges based on protein active sites, and Autodock Vina 1.1.2 software was used for molecular docking. PyMOL was used to show the docking diagram of the target protein and the active component. Based on the principle of lowest binding free energy, the key active components that showed the best docking with each key target protein were determined. This lower binding free energy led to more stable binding between the target protein and the active component, resulting in a higher possibility of interaction between the two components. The design flow chart of this study is shown in Figure 1.
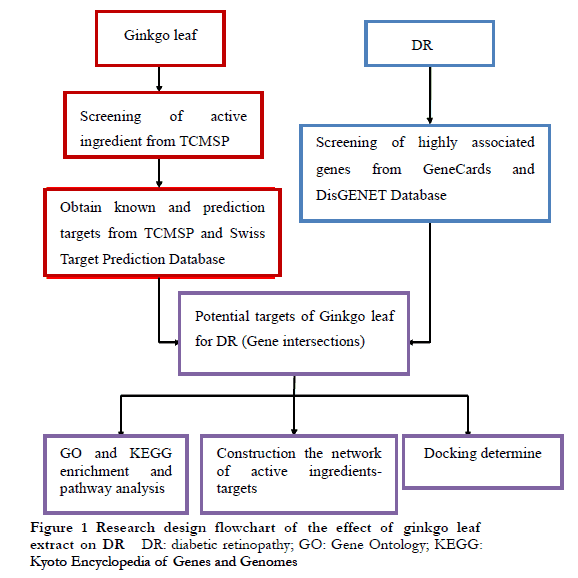
2 Results
2.1 Active components in ginkgo leaf extracts
A total of 307 Ginkgo leaf compounds were retrieved from the TCMSP database. By setting the screening conditions of oral bioavailability ≥ 30% and drug-likeness ≥ 0.18, Ginkgolide A, a component in ginkgo leaf extract with high content and pharmacological activity, was obtained from the supplementary literature 18, and a total of 27 active components in ginkgo leaf extract were obtained. Basic information of active components in ginkgo leaf extract is shown in Table 1.
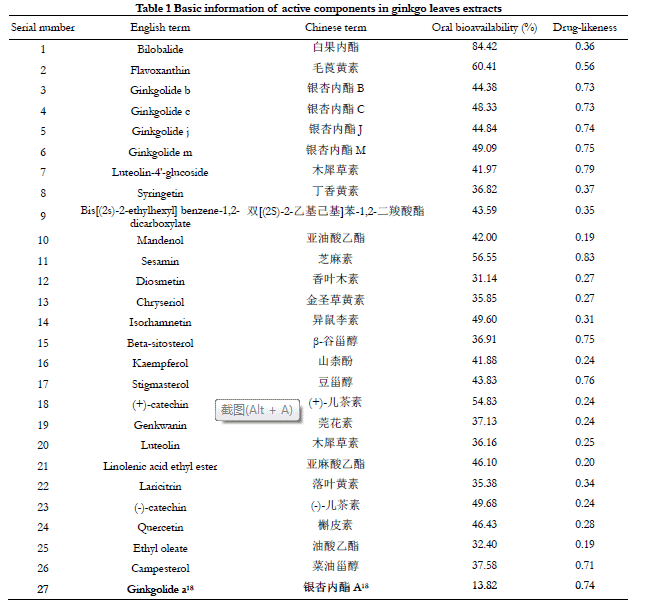
2.2 Potential therapeutic targets of ginkgo leaf extracts against DR
A total of 312 effective targets of 13 active components were obtained after combining the 118 targets obtained from the Swiss target prediction database and 224 targets from the TCMSP database. Another 196 highly related targets were identified from 1,553 DR-related therapeutic targets retrieved from the GeneCards and DisGENET databases. By intersecting the 312 targets of active components in ginkgo leaf extracts and the 196 DR-related therapeutic targets, a total of 34 potential therapeutic targets of ginkgo leaf extract against DR were identified (Figure 2).

2.3 Functional enrichment analysis and pathway analysis
2.3.1 Biological functions of potential therapeutic targets of ginkgo leaves against DR
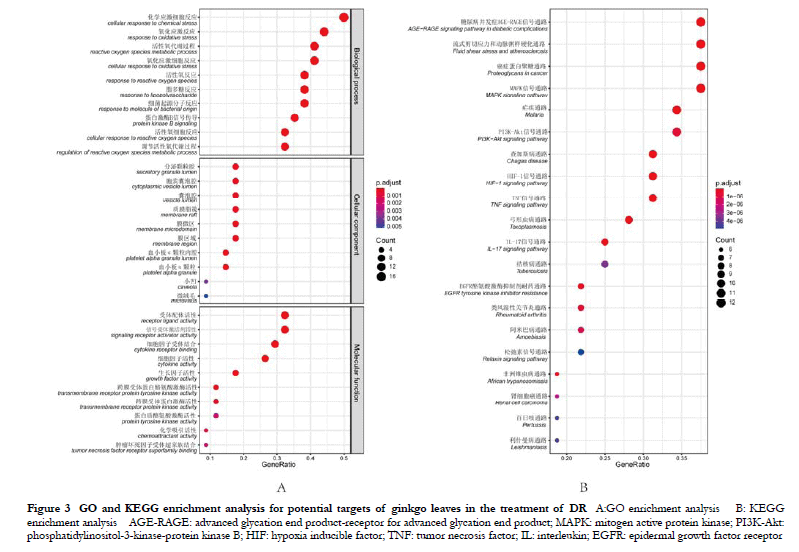
The targets were mainly involved in the processes of chemical stress, oxidative stress, reactive oxygen species metabolism, and protein kinase B signaling. Among the cell components, the targets were mainly found in the secretory granule lumen, the cytosolic vesicle lumen, the membrane region, and the lipid raft pathways. Regarding molecular functions, the targets were mainly involved in the processes of receptor ligand activity, signaling receptor activator activity, cytokine receptor binding, cytokine activity, and growth factor activity (Figure 3A).
2.3.2 Biological signaling pathways of potential targets of ginkgo leaf extracts for the treatment of DR
The targets were enriched in various signaling pathways, including the advanced glycation end product-receptor for advanced glycation end product (AGE-RAGE) signaling pathway, hemodynamics-related flow shear stress and atherosclerotic pathway, proliferation and regulation of angiogenesis related mitogen active protein kinase (MAPK) signaling pathway, phosphatidylinositol-3-kinase-protein kinase B (PI3K-Akt) signaling pathway, hypoxia stress-related hypoxia inducible factor-1 (HIF-1) signaling pathway, immunity and inflammation-related tumor necrosis factor (TNF) signaling pathway, and interleukin, IL)-17 signaling pathway (Figure 3B).
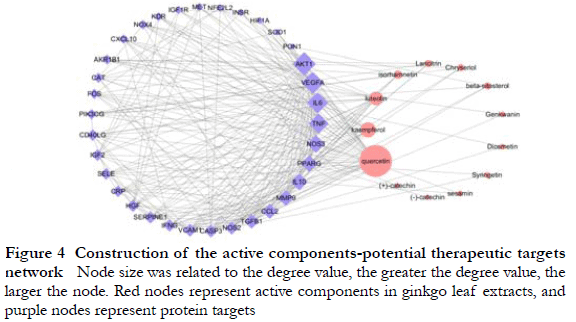
2.4 The active components-potential therapeutic targets network of ginkgo leaf extract and its key nodes
The top four key active components with higher degrees in the network were quercetin, kaempferol, luteolin, and isorhamnetin, and the top eight key targets were serine/threonine protein kinase 1, vascular endothelial growth factor A, IL-6, TNF, nitric oxide synthase 3, peroxisome proliferator activated receptor-γ, IL-10, and matrix metalloproteinase-9 (Figure 4).

2.5 Validation of binding effects between key active components and key targets in ginkgo leaf extracts
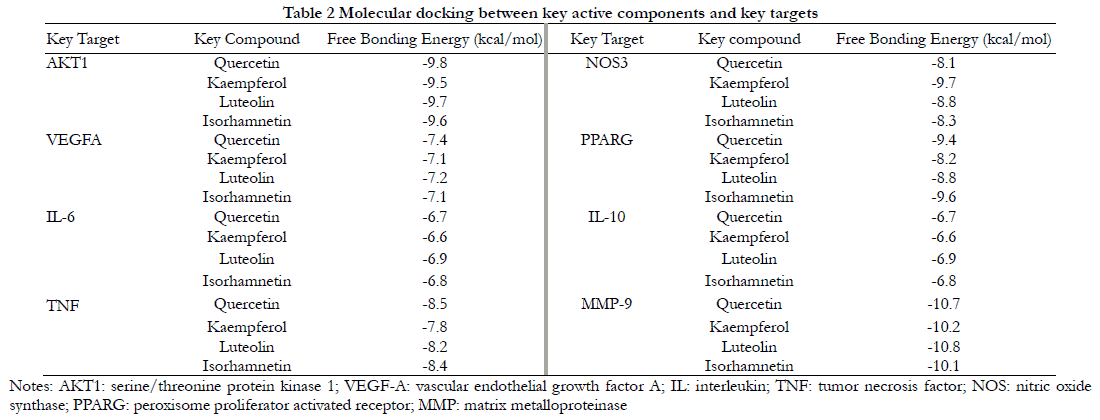
The docking results of key active components with key targets showed that the binding free energies were all less than or equal to -5.0 kcal/mol, indicating that the key active components had good binding with key targets (Table 2). Specifically, quercetin was connected with hydrogen bonds of Ser205, Thr211, Ile290, and Lys268 of AKT1 (Figure 5A), Leu66 of VEGFA (Figure 5B), and Tyr123, Ser107, Gln110, and Glu124 of TNF (Figure 5C). Kaempferol was docked by hydrogen bounding via Met358 of NOS3 (Figure 5D). Luteolin was hydrogen-bonded to Arg16, Leu19, and Arg24 and Gln28 of IL-6 (Figure 5E), Glu142 of IL-10 (Figure 5F), and Met247, Ala189, and Gln227 of MMP-9 (Figure 5G). Isorhamnetin was hydrogen-bonded to Gln273 and Thr268 of PARRG (Figure 5H).

3 Discussion
In this study, a total of 27 active compounds of ginkgo leaf extract and 34 potential therapeutic targets against DR were identified. According to the degree values of the interaction network nodes between the active components of ginkgo leaf extract and their potential therapeutic targets, four key active components were identified, including quercetin, kaempferol, luteolin, and isorhamnetin, and eight key therapeutic targets of AKT1, VEGF-A, IL-6, TNF, NOS3, PPARG, IL-10, and MMP-9. Previous studies have indicated that these key active components are essential for the treatment of DR. For example, quercetin reduced the production of TNF and IL-6 induced by high glucose, and the expression of MMP-9 and VEGF in rats with DR 19-20. Luteolin and quercetin inhibit the VEGF-induced neovascularization, both in vitro and in vivo 21–23. Kaempferol inhibits the angiogenesis of human retinal endothelial cells in a high glucose environment through the targeted inhibition of VEGF 24-25. Additionally, the results of molecular docking between key active components and key targets also verified the binding activity between them.
In this study, 34 potential targets of Ginkgo leaves for the treatment of DR were subjected to GO and KEGG enrichment analyses. The results showed that active components in ginkgo leaves extracts regulated AGE-RAGE, MAPK, PI3K-Akt, HIF-1, TNF, and IL-17 signal pathways in lipid rafts, secretory granule cavities, cytoplasmic vesicle cavities, and other cell components. This meant that it could delay DR progress by reducing inflammation, antioxidant stress, hypoxia induced injury, and by inhibiting angiogenesis. These signaling pathways may be potential therapeutic targets against DR. Among others, an abnormal AGE-RAGE signaling pathway may cause macrovascular and microvascular complications in diabetes mellitus 26. Binding of AGE to RAGE induces a strong signaling cascade and abnormal activation of downstream signaling pathways such as MAPK and PI3K-AKT. This causes retinal microvascular dysfunctions, such as endothelial dysfunction, pericyte apoptosis, neovascularization, and vascular inflammation 27–29. According to previous studies, hypoxia can induce the progression of DR by promoting neovascularization and vascular dystrophy 30-31. In DR, the hypoxia-related HIF-1 signaling pathway regulates retinal neovascularization by affecting VEGF expression, participates in inflammation by regulating the expression of pro-inflammatory factors IL-6 and TNF-α 32-33, and ameliorates retinal microvascular and neuronal damage by inhibiting inflammation mediated by the TNF and IL-17 signaling pathways 33-34. These results suggested that ginkgo leaf extract played a role in the treatment of DR by acting on multicellular types, regulating multiple signaling pathways, and producing different biological functions.
In conclusion, the active components, key targets, and underlying mechanism of ginkgo leaf extract in the treatment of DR were analyzed by network pharmacology, and the binding activity between compounds and the targets were verified by molecular docking. Together, this study provided the basis for further understanding of the pharmacological mechanism of ginkgo leaves extract in the treatment of DR. However, the results need to be further verified by experimental studies.
Conflict of Interest All authors declare that there is no conflict of interest.
Acknowledgments Thanks to Professor Yan Biao of Eye & ENT Hospital of Fudan University for his help.
Author Contributions Bai Wen: conducted experiments, data collation, paper writing and revision; Jiang Qin: experimental design, experimental guidance, and review of the manuscript draft.
References
[1] Wong TY, Cheung CM, Larsen M, et al. Diabetic retinopathy[J/OL]. Nat Rev Dis Primers, 2016, 2:16012[2020-12-28]. http://www.ncbi.nlm.nih.gov/pubmed/27159 554. DOI: 10.1038/nrdp.2016.12.
[2] Campochiaro PA, Akhlaq A. Sustained suppression of VEGF for treatment of retinal/choroidal vascular diseases[J/OL]. Prog Retin Eye Res, 2021, 83:100921[2020-12-28]. http://www.ncbi.nlm.nih.gov/pubmed/33248215. DOI: 10.1016/j.preteyeres.2020.100921.
[3] American Diabetes Association. 10. Microvascular complications and foot care: standards of medical care in diabetes-2018[J]. Diabetes Care, 2018, 41(Suppl 1):S105-S118. DOI: 10.2337/dc18-S010.
[4] Wong TY, Cheung CM, Larsen M, et al. Exploring the potential of traditional herbs in the management of diabetic retinopathy: an overview[J]. Drug Res (Stuttg), 2020, 70(7):298-309. DOI: 10.1055/a-1148-3950.
[5] Chinese Association of Integrative Medicine, Chinese Medical Doctor Association, National Clinical Research Center for Chinese Medicine Cardiology, et al. Chinese expert consensus on clinical application of oral Ginkgo biloba preparations (2020)[J]. Chin J Integr Med, 2021, 27(3):163-169. DOI: 10.1007/s11655-021-3289-6.
[6] Bucolo C, Marrazzo G, Platania CB, et al. Fortified extract of red berry, Ginkgo biloba, and white willow bark in experimental early diabetic retinopathy[J]. J Diabetes Res, 2013, 2013:432695[2020-12-29]. http://www.ncbi.nlm.nih.gov/pubmed/23762874. DOI: 10.1155/2013/432695.
[7] Zhu CY, Yi Q, Ma JL, et al. Clinical evaluation of ginkgo biloba extract for diabetic retinopathy[J]. Int Eye Sci, 2016, 16(2):361-364. DOI: 10.3980/j.issn.1672-5123.2016.2.45.
[8] Jiang R, Zheng YZ, Ren BY, et al. Effect of ginkgo leaf extract on the neural protection of diabetic retinopathy[J]. Int Eye Sci, 2015, 15(8):1327-1331. DOI: 10.3980/j.issn.1672-5123.2015.8.06.
[9] Sun YY, Yang X, An HJ, et al. Protective effect of extract of ginkgo biloba on oxidative damage of human Müller cells [J]. Chin J Exp Ophthalmol, 2015, 33(9):805-810. DOI: 10.3760/cma.j.issn.2095-0160.2015.09.008.
[10] Strømgaard K, Nakanishi K. Chemistry and biology of terpene trilactones from Ginkgo biloba[J]. Angew Chem Int Ed Engl, 2004, 43(13):1640-1658. DOI: 10.1002/anie.200300601.
[11] Ilieva I, Ohgami K, Shiratori K, et al. The effects of Ginkgo biloba extract on lipopolysaccharide-induced inflammation in vitro and in vivo[J]. Exp Eye Res, 2004, 79(2):181-187. DOI: 10.1016/j.exer.2004.03.009.
[12] Lv Z, Shan X, Tu Q, et al. Ginkgolide B treatment regulated intestinal flora to improve high-fat diet induced atherosclerosis in ApoE-/- mice[J/OL]. Biomed Pharmacother, 2021, 134:111100[2020-12-29]. http://www.ncbi.nlm.nih.gov/pubmed/33341056. DOI: 10.1016/j.biopha.2020.111100.
[13] Liu XW, Yang JL, Niu W, et al. Human pharmacokinetics of ginkgo terpene lactones and impact of carboxylation in blood on their platelet-activating factor antagonistic activity[J]. Acta Pharmacol Sin, 2018, 39(12):1935-1946. DOI: 10.1038/s41401-018-0086-7.
[14] Zhang C, Zu J, Shi H, et al. The effect of Ginkgo biloba extract (EGb 761) on hepatic sinusoidal endothelial cells and hepatic microcirculation in CCl4 rats[J]. Am J Chin Med, 2004, 32(1):21-31. DOI: 10.1142/S0192415X04001692.
[15] Jiao X, Jin X, Ma Y, et al. A comprehensive application: molecular docking and network pharmacology for the prediction of bioactive constituents and elucidation of mechanisms of action in component-based Chinese medicine[J/OL]. Comput Biol Chem, 2021, 90:107402[2020-12-29]. http://www.ncbi.nlm.nih.gov/pubmed/3333 8839. DOI: 10.1016/j.compbiolchem.2020.107402.
[16] Chen Y, Xing Q, Huang ZR. Network pharmacology analysis of metformin against diabetic retinopathy[J]. Chin J Exp Ophthalmol, 2020, 38(12):1011-1015. DOI: 10.3760/cma.j.cn115989-20191215-00543.
[17] Liu JP, Li XR. Application of network pharmacology in ophthalmology[J]. Chin J Exp Ophthalmol, 2020, 38(12):1083-1086. DOI: 10.3760/cma.j.cn115989-20200224-00104.
[18] Sarkar C, Quispe C, Jamaddar S, et al. Therapeutic promises of ginkgolide A: a literature-based review[J/OL]. Biomed Pharmacother, 2020, 132:110908[2020-12-29]. http://www.ncbi.nlm.nih.gov/pubmed/33254431. DOI: 10.1016/j.biopha.2020. 110908.
[19] Chen B, He T, Xing Y, et al. Effects of quercetin on the expression of MCP-1, MMP-9 and VEGF in rats with diabetic retinopathy[J]. Exp Ther Med, 2017, 14(6):6022-6026. DOI: 10.3892/etm.2017.5275.
[20] Dokumacioglu E, Iskender H, Sen TM, et al. The effects of hesperidin and quercetin on serum tumor necrosis factor-alpha and interleukin-6 levels in streptozotocin-induced diabetes model[J]. Pharmacogn Mag, 2018, 14(54):167-173. DOI: 10.4103/pm.pm_41_17.
[21] Lupo G, Cambria MT, Olivieri M, et al. Anti-angiogenic effect of quercetin and its 8-methyl pentamethyl ether derivative in human microvascular endothelial cells[J]. J Cell Mol Med, 2019, 23(10):6565-6577. DOI: 10.1111/jcmm.14455.
[22] Li F, Bai Y, Zhao M, et al. Quercetin inhibits vascular endothelial growth factor-induced choroidal and retinal angiogenesis in vitro[J]. Ophthalmic Res, 2015, 53(3):109-116. DOI: 10.1159/000369824.
[23] Bagli E, Stefaniotou M, Morbidelli L, et al. Luteolin inhibits vascular endothelial growth factor-induced angiogenesis; inhibition of endothelial cell survival and proliferation by targeting phosphatidylinositol 3′-kinase activity[J]. Cancer Res, 2004, 64(21):7936-7946. DOI: 10.1158/0008-5472.CAN-03-3104.
[24] Xu XH, Zhao C, Peng Q, et al. Kaempferol inhibited VEGF and PGF expression and in vitro angiogenesis of HRECs under diabetic-like environment[J/OL]. Braz J Med Biol Res, 2017, 50(3):e5396[202-01-03]. http://www.ncbi.nlm.nih.gov/pubmed/28273207. DOI: 10.1590/1414-431X20165396.
[25] Chin HK, Horng CT, Liu YS, et al. Kaempferol inhibits angiogenic ability by targeting VEGF receptor-2 and downregulating the PI3K/AKT, MEK and ERK pathways in VEGF-stimulated human umbilical vein endothelial cells[J]. Oncol Rep, 2018, 39(5):2351-2357. DOI: 10.3892/or.2018.6312.
[26] Ramasamy R, Yan SF, Schmidt AM. Receptor for aGE (RAGE): signaling mechanisms in the pathogenesis of diabetes and its complications[J]. Ann N Y Acad Sci, 2011, 1243:88-102. DOI: 10.1111/j.1749-6632.2011.06320.x.
[27] Xu J, Chen LJ, Yu J, et al. Involvement of advanced glycation end products in the pathogenesis of diabetic retinopathy[J]. Cell Physiol Biochem, 2018, 48(2):705-717. DOI: 10.1159/000491897.
[28] Zong H, Ward M, Stitt AW. AGEs, RAGE, and diabetic retinopathy[J]. Curr Diab Rep, 2011, 11(4):244-252. DOI: 10.1007/s11892-011-0198-7.
[29] Jacot JL, Sherris D. Potential therapeutic roles for inhibition of the PI3K/Akt/mTOR pathway in the pathophysiology of diabetic retinopathy[J/OL]. J Ophthalmol, 2011, 2011:589813[2021-01-04]. http://www.ncbi.nlm.nih.gov/pubmed/22132311. DOI: 10.1155/2011/589813.
[30] Li HY, Yuan Y, Fu YH, et al. Hypoxia-inducible factor-1α: a promising therapeutic target for vasculopathy in diabetic retinopathy[J/OL]. Pharmacol Res, 2020, 159:104924[2021-01-04]. http://www.ncbi.nlm.nih.gov/pubmed/32464323. DOI: 10.1016/j.phrs.2020.104924.
[31] Nyengaard JR, Ido Y, Kilo C, et al. Interactions between hyperglycemia and hypoxia: implications for diabetic retinopathy[J]. Diabetes, 2004, 53(11):2931-2938. DOI: 10.2337/diabetes.53.11.2931.
[32] Wei J, Jiang H, Gao H, et al. Blocking Mammalian Target of rapamycin (mTOR) attenuates HIF-1α pathways engaged-vascular endothelial growth factor (VEGF) in diabetic retinopathy[J]. Cell Physiol Biochem, 2016, 40(6):1570-1577. DOI: 10.1159/000453207.
[33] Gao X, Li Y, Wang H, et al. Inhibition of HIF-1α decreases expression of pro-inflammatory IL-6 and TNF-α in diabetic retinopathy[J/OL]. Acta Ophthalmol, 2017, 95(8):e746-e750[2021-01-04]. http://www.ncbi.nlm.nih.gov/pubmed/27288252. DOI: 10.1111/aos.13096.
[34] Liu S, Lin YU, Liu X. Protective effects of SIRT1 in patients with proliferative diabetic retinopathy via the inhibition of IL-17 expression[J]. Exp Ther Med, 2016, 11(1):257-262. DOI: 10.3892/etm.2015.2877.